Das Leben entschlüsseln
Die Schnittstelle von Biologie & Code
Moderne Medizin ist längst nicht mehr nur Chemie - sie ist eine Data-Science-Challenge. Die Bioinformatik bildet hierbei die entscheidende Übersetzungsebene. Sie fusioniert Biologie, Mathematik und Informatik, um aus gewaltigen Datensätzen - von DNA-Sequenzen bis zu Proteinstrukturen - handlungsrelevante medizinische Erkenntnisse zu gewinnen. Sie ist der Schlüssel, um neue Gene zu entdecken, komplexe Krankheiten zu verstehen und präzise, personalisierte Therapien zu entwickeln.
Biologische Datenanalyse
Interpretation genomischer und proteomischer Landschaften, um signifikante biologische Signale vom bloßen "Rauschen" zu trennen.
Algorithmische Effizienz
Einsatz effizienter Algorithmen zum Durchsuchen, Alignieren und Analysieren riesiger Sequenzdatenbanken und molekularer Netzwerke.
Machine Learning & Deep Learning
Aufdeckung verborgener Muster und Entwicklung prädiktiver Modelle zur Entschlüsselung komplexer biologischer Abhängigkeiten.
Skalierbare Infrastruktur
Architektur von High-Performance-Storage- und Computing-Umgebungen zur Bewältigung des biologischen "Big Data".
Von Molekülen zu Maschinen
Innovation in der Bioinformatik vorantreiben
Aufbauend auf meinem Fundament in Bioinformatik und Data Science schlage ich heute als Head of Technology & Innovation bei MOLEQLAR Analytics GmbH die Brücke zwischen Life Sciences und Advanced Computing. Mein Ziel ist es, komplexe biologische Daten in handlungsrelevante Erkenntnisse zu verwandeln.
Meine Expertise erstreckt sich von der massenspektrometrie-basierten Proteomik, Epigenetik und Deep Learning bis hin zu Cloud-Infrastrukturen, Algorithmen-Design und Innovationsstrategie. Durch die Verbindung von wissenschaftlicher Neugier und technologischer Leadership entwickle ich datengetriebene Lösungen, die den Entdeckungsprozess beschleunigen und die Grenzen biotechnologischer Innovation erweitern.
Case Study: Vorhersage von Peptid-Ladungszuständen in der LC-MS
In diesem Projekt widmete ich mich der Herausforderung, die Vorläufer-Ladungszustände (Precursor Charge States) von Peptiden in LC-MS-Experimenten präzise vorherzusagen. Grundlage hierfür bildete der PROSPECT-Datensatz.
Zur Lösung entwickelte ich verschiedene Deep-Learning-Architekturen von Grund auf neu. Der Fokus lag auf Multi-Class-Klassifikationsmodellen, basierend auf Embedding Layers und Dense Networks. Zusätzlich experimentierte ich mit fortgeschrittenen Multi-Head- und Multi-Label-Ansätzen, um die Modellarchitektur optimal an die chemische Komplexität anzupassen.
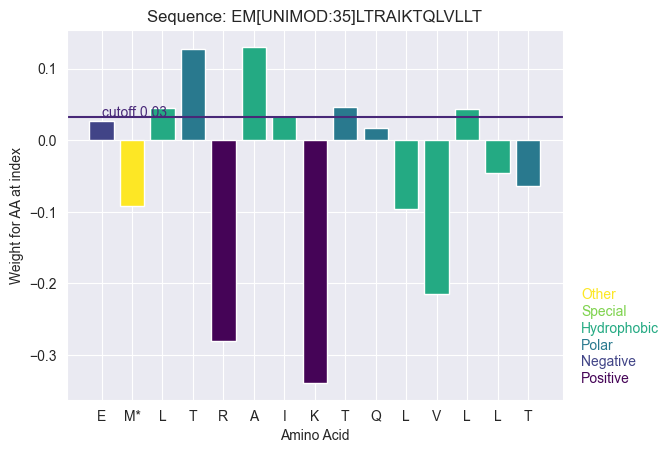
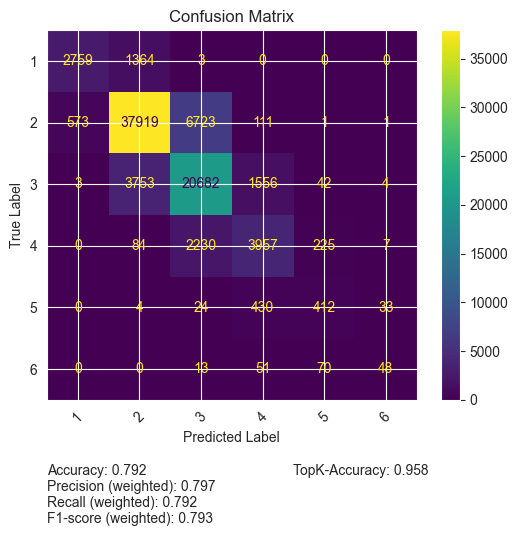
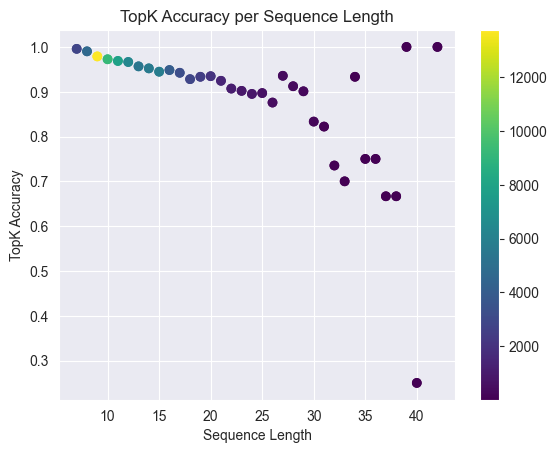
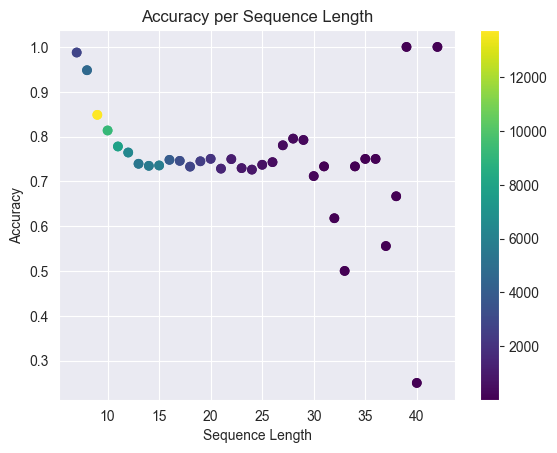
Die folgende Abbildung visualisiert die Vorhersagen für eine exemplarische Proteinsequenz sowie die Performance-Metriken.
Ist das Modell unpräzise? Kann es die Komplexität abbilden? Oder bildet es die Realität ab?
Sind Ladungszustände binär? Oder wird hier vielmehr eine chemische Unsicherheit quantifiziert?
Und jetzt sind wir genau dort: in der Bioinformatik. Im Schnittpunkt von Biochemie, Statistik und Daten.